Zoonotic Bacteriology and Mycoplasmatology
Zoonotic bacteriology:
A broader understanding of health and disease demands a unity of approach achievable only through a consilience of human, domestic animal and wildlife health. Any disease or infection that is naturally transmissible from vertebrate animals to humans and vice-versa is classified as a zoonosis. Our research projects focus on the following zoonotic bacterial agents: Francisella tularensis, the causative agent of the highly contagious zoonotic disease, tularemia - primarily a disease of the orders Lagomorpha and Rodentia; Brucella species, the causative agent of brucellosis - it manifests in abortion in females and in epididymitis, orchitis and inflammation of accessory genital glands in males of different domestic and wild animal species; Coxiella burnetii, the causative agent of Q fever - occurs worldwide and had been associated mostly with late-term abortion and reproductive disorders in wild and domestic ruminants; Chlamydiales species - causing ornithosis in avian species and abortion in mammals; and Borrelia species, the causative agent of Lyme borreliosis - the most prevalent vector-borne human disease in the temperate zone of the Northern Hemisphere.
Mycoplasma research:
Poultry
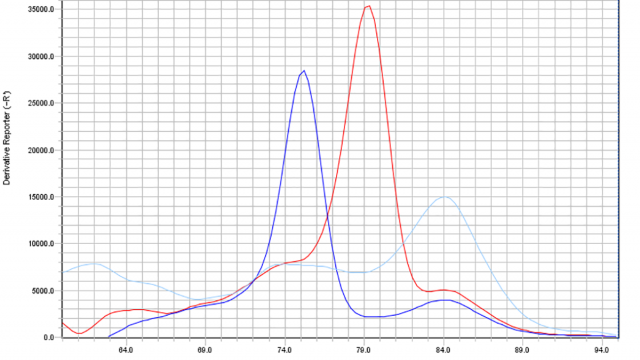
We work in the fields of chicken, turkey (M. gallisepticum, M. synoviae, M. meleagridis, M. iowae) and waterfowl mycoplasmosis (M. anserisalpingitidis, M. anatis, M. anseris, M. cloacale). We study the epidemiology and pathogenesis of poultry mycoplasmosis, as well as participate in the description of novel species (Volokhov et al., 2020). We study the genomics of the different Mycoplasma species (Grózner et al., 2018) and develop methods for molecular epidemiology purposes (Bekő et al., 2019; Kreizinger et al., 2018). We conduct extensive research projects to determine the antibiotic susceptibility profiles (MIC) of the different species in Europe (Kreizinger et al., 2017) and worldwide (Morrow et al., 2020). We are studying the resistance genes and mutations (Bekő et al., 2020) as well as develop molecular assays for the rapid and cost-effective determination of the antibiotic susceptibility profiles of the different species (Bekő et al., 2020). We invent PCR assays for detection (Grózner et al., 2019) and develop DIVA (differentiating infected from vaccinated animals) tests (Sulyok et al., 2019). We intensely work on the development of novel avian Mycoplasma vaccines as well.
Porcine
We are studying M. hyopneumoniae, M. hyorhinis and M. hyosynoviae. We perform genomic research and develop novel genotyping methods (Földi et al., 2020). We determine the antibiotic susceptibility profiles (MIC) of isolates, identify the genetic markers of resistance (Felde et al., 2018) and develop rapid and cost-effective assays for their detection (Felde et al., 2020). In collaboration with pharmaceutical companies we participate in vaccine development projects.
Ruminant
Our research projects are primarily focusing on M. bovis but we examine small ruminant pathogen Mycoplasma species as well. We study the genetics of M. bovis (Sulyok et al., 2014), determine the antibiotic sensitivity of isolates (Sulyok et al., 2014) as well as identify the mutations responsible for antibiotic resistance (Sulyok et al., 2017) and develop diagnostic molecular assays for the rapid detection of these mutations (Sulyok et al., 2018).
Services:
We take part in diagnostic activities, through which the farmers may acquire useful information on diseases threatening their herds and flocks as well as on the possibilities of prevention.
The team has an industrial research and development agreement with different pharmaceutical companies.