Respiratory Bacteriology
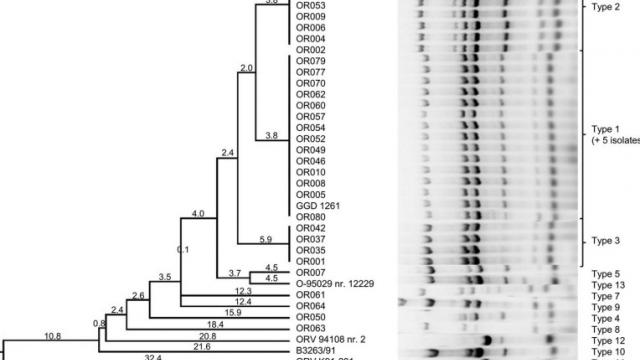
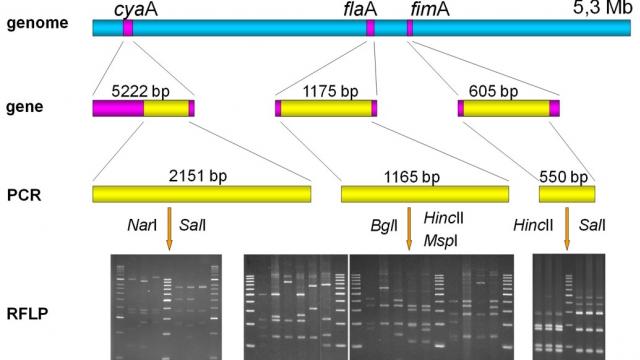
The team studies virulence factors and signs of host adaptation of Bordetella bronchiseptica, a pathogen causing a wide variety of disease in multiple animal hosts (atrophic rhinitis in swine, kennel cough in dogs, bronchopneumonia in cats and rabbits, and sporadic cases in humans) by traditional microbiological and molecular genetic (PCR, PCR-RFLP, sequence and phylogenetic analysis) methods.
Diseases caused by Pasteurella multocida (atrophic rhinitis in swine, haemorrhagic septicaemia in cattle, fowl cholera in poultry, pneumonia in various species, and pasteurellosis in rabbits) result in considerable economic losses for animal husbandry. Traditional (biochemical tests, serotyping, antimicrobial resistance testing) and molecular genetic (PCR, sequence and phylogenetic analysis) methods are applied to analyse P. multocida strains from various animal hosts in order to identify virulence factors and learn more about the pathogenesis of these diseases.
We also examine the prevalence of Bordetella avium, Ornithobacterium rhinotracheale and Riemerella anatipestifer in wild birds and commercial poultry farms. Isolates are characterised by analysing their phenotypic properties, genotyping and determination of antimicrobial susceptibility.
The impact of opportunistic pathogen eukaryotic microorganisms on animal and human health shows a growing trend, therefore our research group is also concerned with the investigation of the molecular epidemiology of infections caused by yeasts (Candida albicans, Malassezia pachydermatis). In order to get a deeper insight into the rate of mutations that occur during various evolutionary events and their effects, we aim to study the genome-level characteristics of these pathogens to reveal the molecular mechanisms controlling virulence and antimycotic resistance.
We offer services to perform laboratory and field trials to test veterinary medicines and vaccines in order to develop modern and more effective preventative methods against infectious diseases of veterinary importance.